- Popularity
- Reliability
- Criteria #3
- Criteria #4
- Criteria #5
-
Characteristics
A protein consists of one peptide folded in a particular way, or several peptides folded together. Such peptides are synthesised very rapidly within living cells, but until recently could only be artificially synthesised in very long, slow processes that had poor yields and gave impure products. Solid phase peptide synthesis (SPPS), developed by R. B. Merrifield, was a major breakthrough allowing for the chemical synthesis of peptides and small proteins. SPPS results in high yields of pure products and works more quickly than classical synthesis (liquid-phase peptide synthesis, LPPS).
The advantages of this method are very considerable. Through the replacement of a complicated isolation procedure for each intermediate product with a simple washing procedure much time is saved. In addition, it has proven possible to increase the yield in each individual step to 99.5% or better, a result which cannot be attained using conventional synthetic approaches. In the example given above the final overall yield would thus be increased from 0.003% to 61%. Finally, this method is also suitable for automation and automatic peptide synthesizers are now commercially available.
SPPS methodology has brought about a revolution in peptide and protein chemistry and thousands of different peptides have now been synthesized using this approach. In addition, this methodology is a completely new approach to organic synthesis. It has created new possibilities in the research fields of peptide-protein chemistry and nucleic acid chemistry It has greatly stimulated progress in biochemistry, molecular biology, pharmacology and medicine. It is also of great practical importance, both for the development of new drugs and for gene technology.
-
Literature reference
・Merrifield, R. B. J. Am. Chem. Soc. 1963, 85, 2149. DOI:10.1021/ja00897a025
・Fmoc method: (a) Chang, C.-D.; Meienhofer, J. Int. J. Pept. Protein Res. 1978, 11, 246. DOI:10.1111/j.1399-3011.1978.tb02845.x (b) Gongora-Benitez, M.; Tulla-Puche, J.; Albericio, F. ACS Comb. Sci. 2013, 15, 217. DOI: 10.1021/co300153c
・Coin, I.; Beyermann, M.; Bienert, M. Nat. Protoc. 2007, 2, 3247. doi:10.1038/nprot.2007.454
<LPPS>
・Kisfaludy, L.; Schon, I.; XSzirtes, T.; Nyeki, O.; Low, M. Tetrahedron Lett. 1974, 15, 1785. doi:1785.10.1016/S0040-4039(01)82579-X
・ Anderson, L.; Blomberg, L.; Fiegel, M.; Lepsa, L.; Nilsson, B.; Verlander, M. J. Pept. Sci. 2000, 55, 227. [abstract]
・Carpino, L. A.; Ghassemi, S.; Ionescu, D.; Ismail, M.; Sadat-Aalaee, D.; Truran, G.; Mansour, E. M.; Siwruk, G. A.; Eynon, J. S.; Morgan, B. Org. Process Res. Dev. 2003, 7, 28. DOI: 10.1021/op0202179
・Eggen, I. F. Org. Process Res. Dev. 2005, 9, 98. DOI: 10.1021/op049864l
<Review of SPPS>
・ Amblard, M.; Fehrentz, J.-A.; Martinez, J.; Subra, G. Mol. Biotechnol. 2006, 33, 239.
・ Pattabiraman, V. R.; Bode, J. W. Nature 2011, 480, 471. doi:10.1038/nature10702
<General Review of Chemical Synthesis of Peptides/Proteins>
・ Humphrey, J. M.; Chamberlin, A. R. Chem. Rev. 1997, 97, 2243. DOI: 10.1021/cr950005s
・ Bray, B. L. Nat. Rev. Drug Discov. 2003, 2, 587. doi:10.1038/nrd1133
・ Nilsson, B. L.; Soellner, M. B.; Raines, R. T. Annu. Rev. Biophys. Biomol. Struct. 2005, 34, 91. DOI:10.1146/annurev.biophys.34.040204.144700
・ Kent, S. B. H. Chem. Soc. Rev. 2009, 38, 338. DOI: 10.1039/b700141j
・ Pattabiraman, V. R.; Bode, J. W. Nature 2011, 480, 471. doi:10.1038/nature10702
・ Stolzew, S. C.; Kaiser, M. Synthesis 2012, 44, 1755. DOI: 10.1055/s-0031-1289765
-
Reaction mechanism
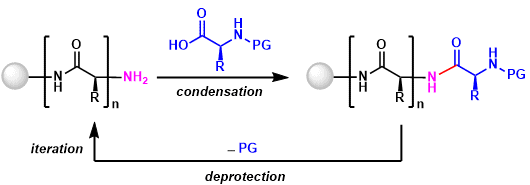
The process consists of five steps carried out in a cyclic fashion.
Step 1 – Attaching an amino acid to the polymer: The amino acid is reacted with a molecule known as a “linkage agent” that enables it to attach to a solid polymer, and the other end of the linkage agent is reacted with the polymer support.
Step 2 – Protection: An amino acid is an acid with a basic group at one end and an acid group at the other. To prevent an amino acid from reacting with itself, one of these groups is reacted with something else to make it unreactive.
Step 3 – Coupling: The protected amino acid is then reacted with the amino acid attached to the polymer to begin building the peptide chain.
Step 4 – Deprotection: The protection group is now removed from the acid at the end of the chain so it can react with the next acid to be added on. The new acid is then protected (Step 2) and the cycle continues until a chain of the required length has been synthesised.
Step 5 – Polymer removal: Once the desired peptide has been made the bond between the first amino acid and the linkage agent is broken to give the free peptide.
-
Example of reactions
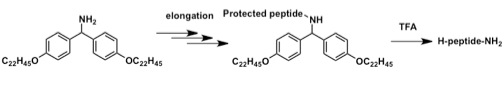
This approach combines the advantages of both SPPS and LPPS. The fluorene-type protecting group exhibits an effect similar to that of the chlorotrityl linker used in SPPS, namely the avoidance of diketopiperazine formation. The selective detachment of this C-terminal protecting group with the protecting groups of the side chain intact leads to production of protected peptide acids that are useful for the convergent synthesis of long-chain peptides.
As a result, this easy to implement and efficient LPPS protocol (AJIPHASE) was successfully extended to the synthesis of peptide amides, which account for the majority of current peptide drugs, and for which a sufficient LPPS method has not yet been developed to date.[1]
-
Bibliography
[1] (a) Takahashi, D.; Yamamoto, T. Tetrahedron Lett. 2012, 53, 1936. doi:10.1016/j.tetlet.2012.02.006(b) Takahashi, D.; Yano, T.; Fukui, T. Org. Lett. 2012, 14, 4515. DOI: 10.1021/ol302002g
-
Related Books
[amazonjs asin=”0199637245″ locale=”US” title=”Fmoc Solid Phase Peptide Synthesis: A Practical Approach (Practical Approach Series)”][amazonjs asin=”0716701448″ locale=”US” title=”Solid Phase Peptide Synthesis”][amazonjs asin=”3527318674″ locale=”US” title=”Peptides: Chemistry and Biology”][amazonjs asin=”047062776X” locale=”US” title=”Oxidation of Amino Acids, Peptides, and Proteins: Kinetics and Mechanism (Wiley Series of Reactive Intermediates in Chemistry and Biology)”]
-
Related Links
・Peptide (Wikipedia) ・Peptide Synthesis (Wikipedia) ・Dicyclohexylcarbodiimide (Wikipedia) ・Carbodiimide (Wikipedia) ・PyBOP (Wikipedia) ・Hydroxybenzotriazole (Wikipedia) ・Condensation Reaction (Wikipedia) ・Steglich Esterification (organic-chemistry.org) ・Ajiphase Peptide Synthesis

Good
Pingback: Native Chemical Ligation (NCL) | 化学空间 Chem-Station