Uno, S. Kamiya, M..; Yoshihara, T.; Sugawara, K.; Okabe, K.; Tarhan, M. C.; Fujita H.; Funatsu. T.; Okada. Y.; Tobita, S.; Urano., Y. Nat. Chem. 2014, ASAP.
DOI:10.1038/nchem.2002
Single-molecule localization microscopy is used to construct super-resolution images, but generally requires prior intense laser irradiation and in some cases additives, such as thiols, to induce on–off switching of fluorophores. These requirements limit the potential applications of this methodology. Here, we report a first-in-class spontaneously blinking fluorophore based on an intramolecular spirocyclization reaction. Optimization of the intramolecular nucleophile and rhodamine-based fluorophore (electrophile) provide a suitable lifetime for the fluorescent open form, and equilibrium between the open form and the non-fluorescent closed form. We show that this spontaneously blinking fluorophore is suitable for single-molecule localization microscopy imaging deep inside cells and for tracking the motion of structures in living cells. We further demonstrate the advantages of this fluorophore over existing methodologies by applying it to nuclear pore structures located far above the coverslip with a spinning-disk confocal microscope and for repetitive time-lapse super-resolution imaging of microtubules in live cells for up to 1 h.
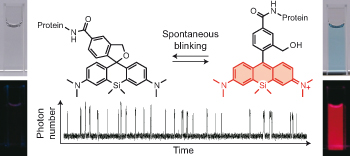
Enabling fluorescence dyes to be fluorescence switchable is essential for acquisition of images beyond the visible light diffraction limit: Super resolution image. dSTORM (direct Stochastic Optical Reconstruction Microscopy)[1] is the one of the most practical methodology because it takes advantage of fluorescence blinking, one of the intrinsic fluorescence switch mechanism. In this paper, the repetitive fluorescence blinking was successfully achieved utilizing fluorophore based on an thermal equilibrium of intramolecular spirocyclization reaction. For the practical application for the super resolution imaging, structural optimization, evaluation of and their basic photo physical properties and actually live imaging were demonstrated.
Structural optimization.
For practical use, following two factors need to be considered. First, most of fluorophores should be dark state(Closed form) leaving only a handful of them in bright state at pH=7.6 evironment which is physiological condition for living cell. Second, the fluorescent state has to last long enough to be detected for the precise acquisition of photon. To fulfill these criteria, pH titillation and laser flash photolysis (LFP) were performed as following.
From these experiment, HMSiR was selected as the most prominent candidate for further experiments because HMSiR not only fulfilled all these requirements but has strong photoresistance, relatively long emission wavelength and high fluorescence quantum yield.[2]
Fluorescence switch
Next, the fluorescence blinking at single molecular level was characterized and compared with Alexa647 and 2MeSiR. As shown below, ony HMSiR exhibited clear fluorescence blinking. This result revealed that the fluorescence blinking is not caused by ROX system (typical dSTORM method) but by thermal equilibrium of intramolecular spirocyclization reaction.
Super resolution image
<In fixed cell>
For comparison, the image obtained from typical microscopy and that of super resolution technic shown below. The left top and left below hand are images taken by TIRFM (Total Internal Reflection Fluorescence) and spinning-disk confocal microscopy respectively. The right images are their corresponding super resolution images.
<Live imaging>
Live imaging was also recorded using dSTORM method. While the average image was blurred due to the diffraction of visible light, the resolution of image in right hand is highly improved with the value of FWHM 83.1 nm.
- Reference
[1] “Subdiffraction-Resolution Fluorescence Imaging with Conventional Fluorescent Probes”
Heilemann, M.; van de Linde, S.; Schüttpelz, M.; Kasper, R.; Seefeldt, B.; Mukherjee, A.; Tinnefeld, P.; Sauer, M. Angew. Chem. Int. Ed. 2008, 47, 6172. DOI: 10.1002/anie.200802376
Eagle eyes: dSTORM uses conventional photoswitchable fluorescent dyes that can be reversibly cycled between a fluorescent and a dark state by irradiation with light of different wavelengths (see picture). This elegant approach can visualize cellular structures with a resolution of approximately 20 nm, far beyond the diffraction limit of light, without the need of an activator molecule.
[2] “A near-infrared fluorophore for live-cell super-resolution microscopy of cellular proteins”
Lukinavičius, G.; Umezawa, K.; Olivier, N.; Honigmann, A.; Yang, G.; Plass, T.; Mueller, V.; Reymond, L.; Corrêa, I. R., Jr; Luo, Z.-G.; Schultz, C.; Lemke, E. A.; Heppenstall, P.; Eggeling, C.; Manley, S.; Johnsson, K. Nat. Chem. 2013, 5, 132. DOI:10.1038/nchem.1546
The ideal fluorescent probe for bioimaging is bright, absorbs at long wavelengths and can be implemented flexibly in living cells and in vivo. However, the design of synthetic fluorophores that combine all of these properties has proved to be extremely difficult. Here, we introduce a biocompatible near-infrared silicon–rhodamine probe that can be coupled specifically to proteins using different labelling techniques. Importantly, its high permeability and fluorogenic character permit the imaging of proteins in living cells and tissues, and its brightness and photostability make it ideally suited for live-cell super-resolution microscopy. The excellent spectroscopic properties of the probe combined with its ease of use in live-cell applications make it a powerful new tool for bioimaging.