Suzuki, Y.;;Endo, M.; Katsuda, Y.; Ou, K.; Hidaka, K.; Sugiyama, H. J.Am. Chem. Soc. 2013, ASAP.
DOI:10.1021/ja408656y
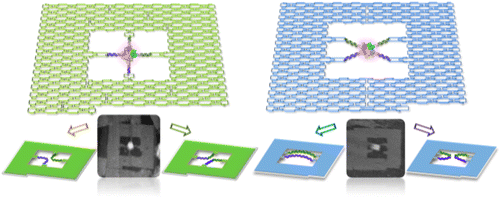
Site-specific recombination involves reciprocal exchange between defined DNA sites. The reaction initiates from the formation of a recombinase–DNA synaptic complex, in which two recombination sites arrange in an appropriate configuration. However, there is incomplete information about how the topological state of the substrate influences the synapsis and outcome of the reaction. Here, we show that Cre-mediated recombination can be regulated by controlling the orientation and topology of the loxP substrate in a DNA frame nanoscaffold. High-speed atomic force microscopy analyses revealed that the loxP-containing substrate strands in the antiparallel orientation can be recombined only through formation of synaptic complexes. By tethering Holliday junction (HJ) intermediates to DNA frames in different connection patterns and using them as a starting substrate, we found that the topological state of the HJ intermediates dictates the outcome of the resolution. Our approach should provide a new platform for structural–functional studies of various DNA targeting enzymes, especially which require formation of synaptic complexes.
DNA is one of the most promising molecules for preparing various self-assembled components and large scaffolds for the production of complicated patterns, and for placing and arranging functional molecules and nanomaterials. Sugiyama and Endo group used a frame-like DNA origami scaff old (DNA frame)17−22 in which two loxP site-containing double-stranded DNAs (dsDNAs) can be arranged in parallel. By combining this system with high-speed atomic force microscopy (HS-AFM) imaging, They directly monitored the Cre-mediated site-specifi c recombination process at a single molecule level in the context of the substrate loxP site orientation that infl uences the synapsis and the subsequent reaction. or antiparallel orientations.
- Related links
Sugiyama group
- Related Books
[amazonjs asin=”3642360769″ locale=”US” title=”DNA Nanotechnology: From Structure to Function”][amazonjs asin=”3642183042″ locale=”US” title=”DNA Computing and Molecular Programming: 16th International Conference, DNA 16, Hong Kong, China, June 14-17, 2010, Revised Selected Papers (Lecture … Computer Science and General Issues)”]